Discovery of a Novel Type of Plant Lnc-eRNA XSER1: Promotes Chromatin Looping to Regulate Broad-Spectrum Disease Resistance in Rice
Enhancers are DNA elements that activate gene expression in response to specific signals. Active enhancers produce bidirectional non-coding RNAs called eRNAs and show H3K27ac histone marks. In mammals, some long non-coding RNAs initiate from enhancers and are known as lnc-eRNAs, they are more stable and linked to tissue-specific regulation. Whether plants possess lnc-eRNAs and whether they play a role in disease defense remains unknown.

On March 19, 2026, Professor Yueqin Chen's team from our School of Life Sciences published a research article online in Molecular Cell entitled “The XSER1 lncRNA acts as an enhancer RNA and confers resistance to pathogens by promoting chromatin loop formation in rice”. This study systematically identifies a class of pathogen-induced lnc-eRNAs XSERs in rice for the first time, and reveals the molecular mechanism of XSER1 that negatively regulates rice disease resistance by recruiting the transcription factor OsbHLH94 to promote chromatin loop formation, thereby activating the expression of the adjacent histone demethylase gene OsJMJ707.
Based on previous discovery that numerous lncRNAs accumulate abnormally upon pathogen infection in rice, the research team predicted enhancers based on DNase I hypersensitive sites (DHS) and combined this with pathogen-responsive lncRNA expression profiles. From 567 lncRNAs responsive to Xanthomonas oryzae pv. oryzae, they screened 64 candidate Lnc-eRNAs, designated XSER (Xanthomonas Susceptible Enhancer RNA). Among these, XSER1 exhibited typical eRNA characteristics, showing rapid and transient upregulation upon pathogen induction. Functional validation demonstrated that both XSER1-RNAi knockdown and xser1-KO (large-fragment deletion) plants displayed enhanced resistance to bacterial blight, bacterial leaf streak, rice blast, and sheath blight diseases, with significantly reduced lesion lengths. These findings establish XSER1 as a disease-susceptibility-associated Lnc-eRNA whose suppression confers broad-spectrum disease resistance in rice.
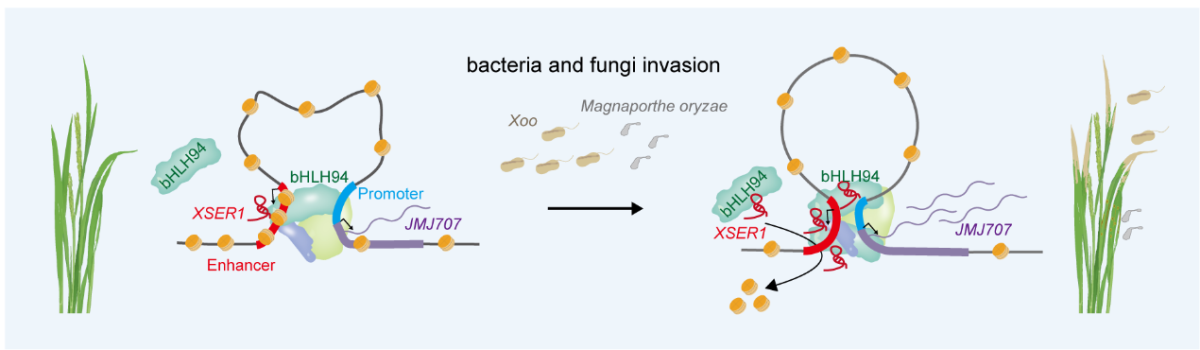
Moreover, the study reveals that, when pathogens attack rice leaves, XSER1 levels increase significantly within hours. The RNA recruits a transcription factor OsbHLH94, to promote chromatin looping and activate the adjacent H3K4 demethy lase gene OsJMJ707, thereby conferring broad-spectrum disease resistance in rice Manipulating XSER1 expression specifically regulates OsJMJ707 transcription without affecting other neigh boring genes, and genetic epistasis places OsJMJ707 downstream of XSER1 in immune regulation, establishing a direct regulatory hierarchy.
It was worthy mentioning that, XSER1 is expressed at extremely low levels under normal conditions, and its knockout does not affect growth and development while significantly enhancing broad-spectrum resistance, breaking the traditional trade-off between disease resistance and growth, and providing a precise target for crop disease-resistance breeding.
AUTHOR CONTRIBUTIONS
C.Y. and J.J. conceived the project, designed and analyzed data, performed experiments, and wrote the manuscript; Y.C., J.-P.L. performed experiments and analyzed data; Y.-H.L., Y.Y., W.-L.Z., W.-T.M., J.-B.Y., Y.-C.Q., Z.-T.C., L.Y., M.-Q.L. performed experiments; Y.-C.Z. performed data analysis. R.-R.H., Y.-F.Z. help supervise the project and designed experiments. Y.-Q.C. supervised the project, designed experiments, and wrote the manuscript.
ACKNOWLEDGMENTS
This research was supported by the National Key Research and Development Program, the National Natural Science Foundation of China, and the Natural Science Foundation of Guangdong Province.
Original article: https://www.cell.com/molecular-cell/fulltext/S1097-2765(26)00127-9



